May 13, 2024 by Bradley Voytek
This weekend I had the honor of giving the keynote address at the UC Berkeley Cognitive Science undergraduate commencement ceremony. There were approximately 150 students, along with their family and friends, in attendance at Zellerbach Hall. Fourteen years ago, in the same location, I was awarded my PhD in neuroscience. And, just as then, my PhD advisor, friend, and mentor, Bob Knight was on stage with me. It was a great homecoming.
I’ve given a lot of talks and taught a lot of classes over the years. I’m normally a semi-extemporaneous speaker, meaning I have a clear vision of what I want to say / teach, but that I come up with the exact wording on the fly.
This is the first time I’ve ever written out the script of what I was going to say. For those who are interested, below is the full text of my address (minus some improvising along the way).
Finally, I want to thank all of the students and family who came up to speak with me afterward! It was a genuine pleasure to meet so many people who are beginning their exciting careers. And a huge thank you to Bob Glushko for nominating me as speaker, David Whitney for doing such an amazing job as master of ceremonies, and Terry Regier for supporting the students.
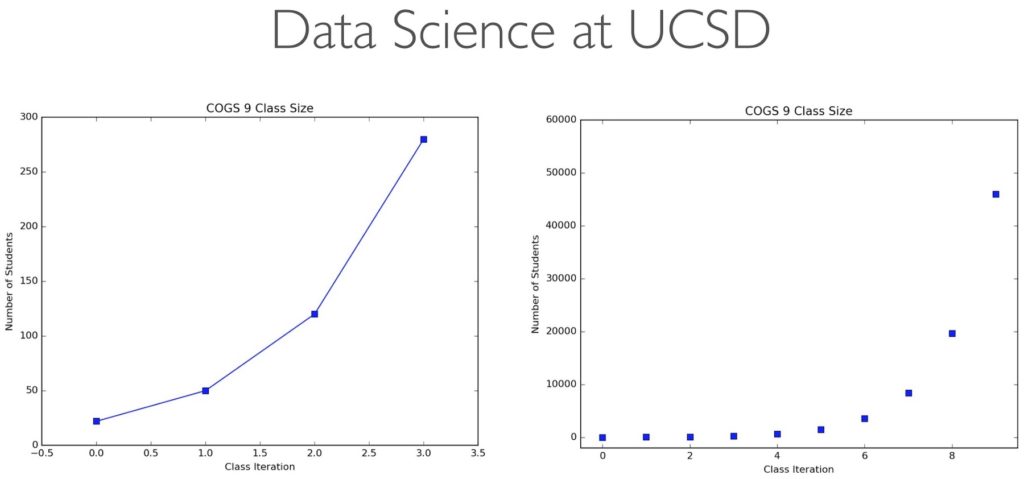
Congratulations, class of 2024! Cognitive Science is different at Berkeley compared to UC San Diego: you don’t have a Cognitive Science department, but rather a distributed program across neuroscience, linguistics, data science, design, computer science, statistics, and so on. That poses unique challenges for you, I’m sure, because it’s such a broad and diverse program.
Of course the biggest challenge is explaining to your parents exactly what Cognitive Science is…
But there is strength in that intellectual diversity! My first real introduction to Cognitive Science was here at Berkeley when I was working on my PhD in neuroscience. I was teaching the MCB 163 neuroanatomy lab. I was initially in WAY over my head – MCB 163 was mostly full of premeds who thought it was funny when the non-biology guy made ridiculous mistakes about basic human physiology. Like the time I confidently proclaimed that humans have two livers… apparently that’s incorrect.
That class also had a lot of students from the CSSA - the Cognitive Science Student Association. As I got to know them better, they began inviting me to their meetings to talk about my research and to ask about general life and academic advice. Which was funny because: A) I wasn’t a Cognitive Scientist, and, B) I for SURE was not a good person to ask academic advice of.
As an undergrad, I began as a physics major… I wanted to study cosmology and astrophysics. But I don’t come from a college family. I was a “whoopsie” baby born to teenaged parents. My dad worked his butt off when I was a kid to make sure I could be where I needed to be to thrive. So the fact that his son wanted to be an astrophysicist was something he delighted in. He loves to tell a story about how, when we were on a road trip between San Diego and Phoenix where I bounced around as a kid, he asked me what I wanted to be. When I told him “astrophysicist” he laughed because he didn’t know what that was.
Even though I don’t come from an “academic” family, I was a great student in high school. So much so that I got accepted to the University of Southern California directly out of my junior year of high school to study astrophysics. I was elated. As part of my scholarship package — USC is an expensive private university outside my family’s reach! – I received work study as a research assistant in a lab studying Bose-Einstein condensates. Don’t worry about what that is… if you don’t know it’s okay. I didn’t either and, well YOU’RE not going to fail out of university for not knowing.
I DID fail out though because WOW was I terrible at physics. I mean really, truly, genuinely terrible. When I went to register for classes at the end of my second year, I was told I’d been on academic probation for too long and was no longer a student there!
But I wasn’t ready to give up. I still loved science, but I had to reassess myself. I basically forced my way back into university by talking to everyone I could, explaining my situation – life was complicated at the time, taking summer classes at community college to prove I could get my grades, and finding people who were willing to listen and help.
This was back in 2000 or so, and several of my closest friends were older than me and had graduated with degrees in computer science. Straight out of undergrad, they landed jobs that paid more than I could have ever imagined.
When I reenrolled, I knew I had to give up physics. I couldn’t switch to computer science, because I wouldn’t be able to graduate within four years. So I switched to Psychology, which I could still complete within the four years, thus minimizing my debt (because by this point I’d lost my scholarship). Psychology was a flexible major, so for my electives I took some programming classes to help fill out my resume and give myself some marketable skills. But, for fun I took courses like Philosophy of Mind, intro to AI programming (in LISP!), sensation and perception… it turns out I accidentally made a Cognitive Science curriculum for myself without even knowing what Cognitive Science was!
I also got another work study job, this time in a neuroscience lab. This lab had decades worth of data that had been collected. A lot of the data were surveys that were digitized in the form of text files. One of the first jobs I was given there was wild: I supposed to open up each file, copy the data out of the file, and paste them into a Excel spreadsheet so that the Postdoc could analyze the results. Based on how many files there were, they estimated it would take two weeks or so.
I HATE this kind of repetitive task and I’m always looking for ways to get the work done without having it be so boring. So I flexed my super crude C++ skills and wrote a quick regex script to pull the data into a CSV. I showed up the next day and was like “here you go”. And it was like I performed some kind of magic trick. They didn’t believe me. So I explained to them how “I wrote some code,” and they just didn’t know what that was.
Magic!
And thus, I became the “tech guy” in the lab, and began digitizing a lot of their processes. I used my knowledge of web development, which I learned from my girlfriend at the time (and wife now!), and coding to help with neuroscience experimental design and data analysis. I realized that this diverse skillset was a huge strength and that, although I wasn’t as good of a programmer as my computer science friends or as good of a researcher as my neuroscience friends, I was better programmer than the neuroscientists and a better neuroscientist than the programmers!
It’s this diverse and distributed nature of Cognitive Science that is its strength. And what an amazing time to be a Cognitive Scientist! The last few years have seen an explosion in deep learning, data science, and AI, which all have their roots in Cognitive Science (many of them at Berkeley and UC San Diego!).
And just like how my crude C++ regex skills seemed like magic to my boss back in 2001, the technologies that are coming out now, that have their roots in Cognitive Science, seem like magic to the rest of the world.
If I’m allowed to revel in my sci-fi and comic book roots for a minute, I’ll quote Arthur C Clarke’s third law: “Any sufficiently advanced technology is indistinguishable from magic.”
And I’ll expand on that and say to all of the parents and guardians here, I’m happy to give you the real explanation of what Cognitive Science is: your students just completed their degrees in magic.
Do we have a sorting hat or how does this work?
It’s silly I know, but I genuinely believe in this idea that we can create technologies that seem like magic. But for me, this metaphor isn’t restricted just to creating technologies. It also applies more broadly to the act of creating new ideas and, with these ideas, shaping our world and ourselves.
After I graduated, I knew I wanted to become a neuroscientist, but I also knew that my GPA was too awful for that to be a realistic shot. So I spent a lot of time talking with my girlfriend (at the time, my wife now) and my closest friends, trying to figure out how to turn my wishful, magical thinking into reality. The first part of the magical spell was to figure out a way to prove that my undergraduate GPA wasn’t an accurate reflection of who I am. Because of my programming skills, I was able to land a job as a full-time researcher running the Positron Emission Tomography scanner at UCLA working for Dr. Edythe London.
This job was awesome. For the most part. What wasn’t exactly made clear to me during the interview was that the radiotracer we used to study brain activity, F18 FDG, though very short lived, is cleared through the bladder. And because we want to minimize the effective radioactive dose to all the participants in our studies, we had to have them use the restroom immediately after the scanning was complete. Buuuut, because some people are… messier… than others, part of my job was to crawl around the bathroom floor with a Geiger counter looking for little radioactive pee pee spots to clean up.
Science!
But I made a name for myself at UCLA’s Brain Mapping Center. Because PET scanning works by injecting a radioactive substance, it requires a nuclear medicine physician to perform the injections. But something like 4-6 months into the job, I was waiting with a participant at the scanner with the rest of the staff, and our physician didn’t show. Finally I went back to my boss’s office, and apparently the guy left a message on her voicemail saying he moved back to New York, that he didn’t want to work there anymore.
So all of these scientists with huge amounts of grant funding were stuck and I was one of the only people who knew how to run the scanner. So I learned that it’s not just a nuclear medicine physician who could perform the injections, but that credentialed nuclear medicine technicians, non-physicians, could also do so. So I called all around LA to find techs who could do this. I proved that I could solve problems. That I could publish research and train others. And in 2004 I was accepted into Berkeley’s new neuroscience PhD program.
When I began my PhD I remember looking at the CVs of researchers whom I admired and seeing page after page after page of amazing awards and scientific achievements. It was daunting. During my very first year here, I even asked my PhD Advisor, and now friend and mentor, Bob Knight, how I got into Berkeley when no one else even gave me an interview (including my home department of UC San Diego Cogsci!). He said – and I’m editing his wonderful bluntness here a bit – “it was obvious you were a… screw up… but a screw up with potential.”
A few years into my PhD, during one of those career discussions with the CSSA students here, someone asked me something like “how is it you’ve been so successful in your research?”
I was like, ARE YOU KIDDING ME?!
But it hit me: hidden behind the plethora of successful outcomes on those CVs I admired were an even greater number of failures. The CSSA students here didn’t know that I was kicked out of undergrad, or that I was immediately placed on academic probation – again! – my first semester as a PhD student here. They didn’t know that Prof. Rich Ivry almost failed me in my qualifying exams. Because we keep all of those things hidden.
So for the last 15 years or so I’ve kept a section at the end of my CV titled, “Rejections and Failures”. I list every grant or fellowship I did not receive, how many times each paper I have published was initially rejected by journal editors.
This was perhaps my most powerful magic spell yet. This act of revealing my otherwise invisible failures has brought into my world some of the most amazing scientists who saw this and said, “I want to work with that guy.”
That’s magic! Taking what was once a real source of shame for me and turning it into a source of strength. It was like a summoning spell for kind, brilliant people who may not have had the most traditional scientific paths. Or, as Bob Knight might call us… “screw ups with potential”. This spell was so powerful that, years later I was even told in private by a very well known senior scientist: “part of the reason the committee decided to give you this award was because we didn’t want our name associated with the failures section of your CV!” I’m still not entirely sure if she was joking or not.
I’ve caught a ton of breaks to help me get to where I am today, and had a lot of excellent mentorship. A lot of it has felt like luck, but I’d be lying if I said it was JUST luck. Because a lot of it was a willful application of my probably naively optimistic view that we can both be good people AND good scientists. That we can push at the boundaries of technology until what we create seems magical, and that we can also summon amazing people to work alongside us to bring about cultural change in science and technology.
So yes “any sufficiently advanced technology is indistinguishable from magic.” I believe that. But I also believe that, “any sufficiently thoughtful application of optimism is indistinguishable from magic.” I have no doubt that your quirky, Berkeley education here in Cognitive Science has armed you with the diversity of skills to create the technologies of the future that will truly feel magical.
And all I can hope is that some of what I’ve said today has resonated with you, and that the stories I’ve told help focus your skills not just toward building amazing magical technologies, but also in building a more optimistic, magical version of yourselves.
]]>